Technology
Tech. 1
NEORNAT’s Strategies of RNA-based & RNA-targeted Therapeutics
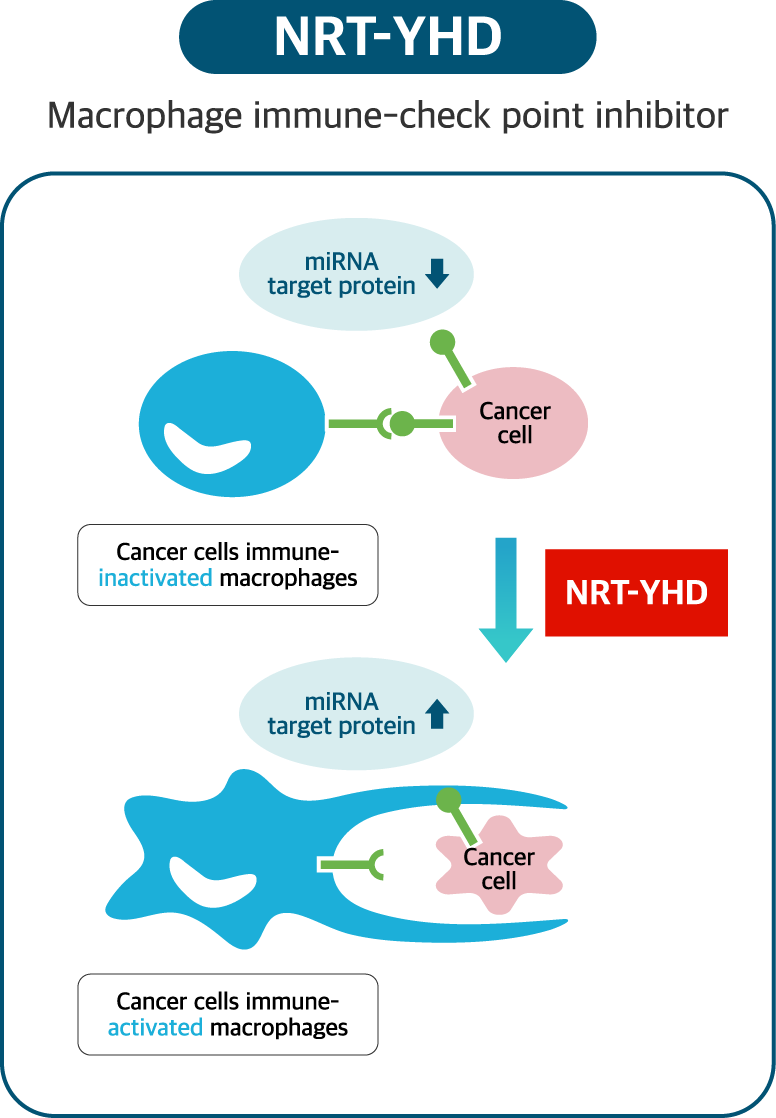
In general, there are several different RNA therapeutic approaches including mRNAs, antisense oligonucleotides (ASOs), siRNAs and miRNAs. miRNA mimics are small double-stranded RNA molecules that associate with and guide the RISC complex to its target mRNA. miRNA inhibitors are small single-stranded RNAs that bind to and suppress their target miRNA, and this results in restored mRNA translation. NEORNAT’s first pipeline, NRT-YHD, is a modified miRNA inhibitor. NEORNAT has discovered novel miRNA which suppresses macrophage phagocytic activation toward cancer cells. NRT-YHD is modified anti-sense miRNA that functions as novel macrophage immune-check point inhibitor.
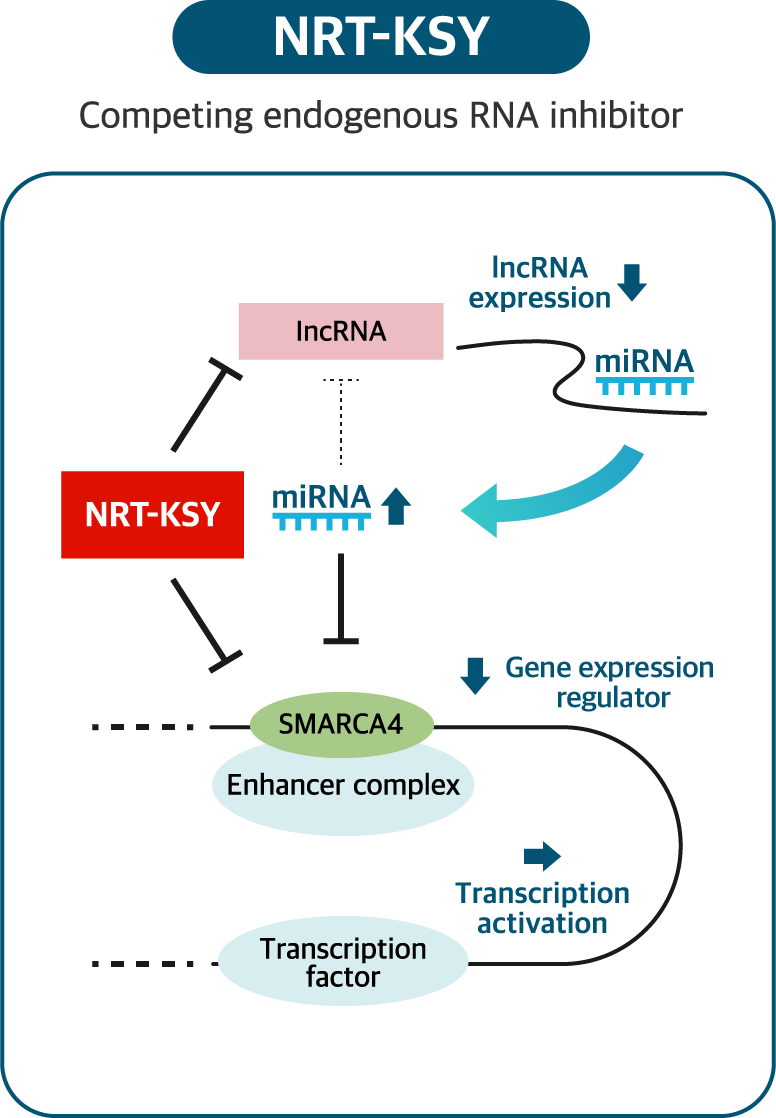
siRNAs are small double-stranded RNA molecules that have exact complementarity to a target mRNA. Once associated with the RISC complex, it binds to its target mRNA and induces gene silencing by preventing translation of the mRNA. NRT-KSY pipeline is double siRNA therapy that target both long non-coding RNA and subunit of SWI/SNF complexes. NEORNAT have demonstrated that these two molecules simultaneously contribute to cancer development and progression. Therefore, it is unique strategy to regulate coding & non-coding RNA for the development of RNA therapeutics in cancer.
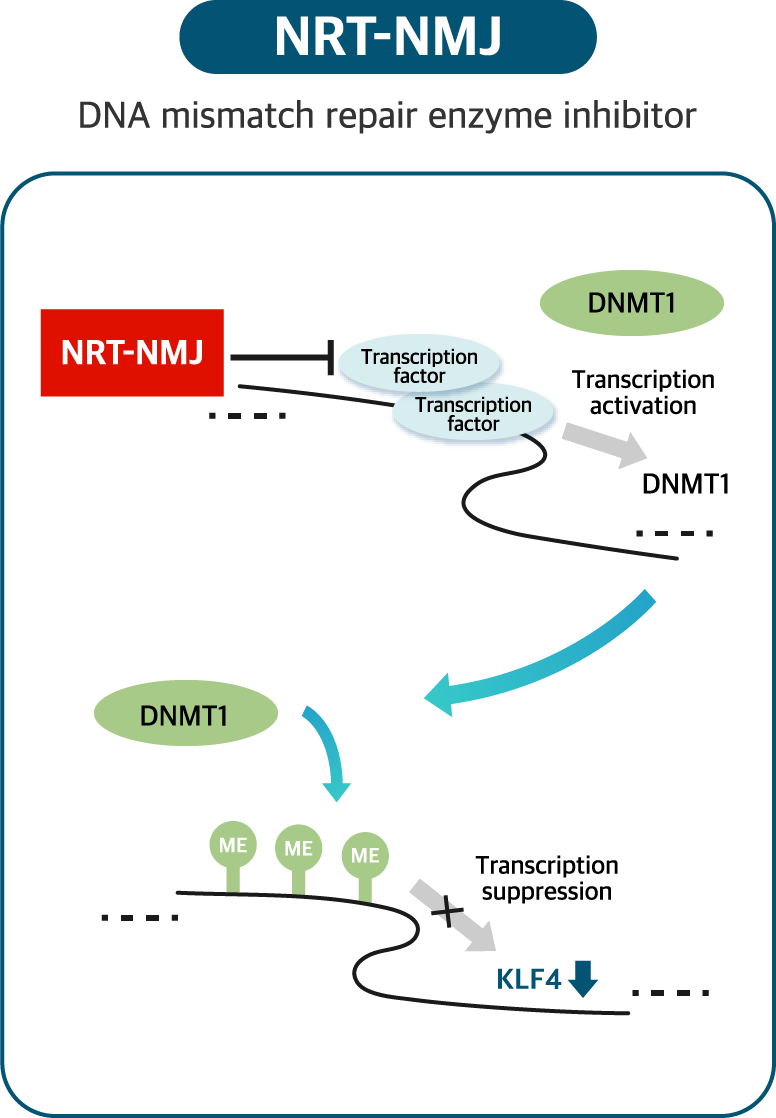
NRT-NMJ pipeline targets novel DNA repair enzyme for cancer treatment. NEORNAT has identified specific DNA repair enzymes which aberrantly deregulated in cancers. Targeting single or combination of these DNA repair enzymes showed strong anti-caner effects in vitro and in vivo.
Tech. 2
In vivo validation system
NEORNAT Inc. has established various cancer models and uses it for research and drug development. In particular, the liver cancer model is a world -class level with a system that can evaluate the effect of liver cancer with real-time without sacrificing animals.
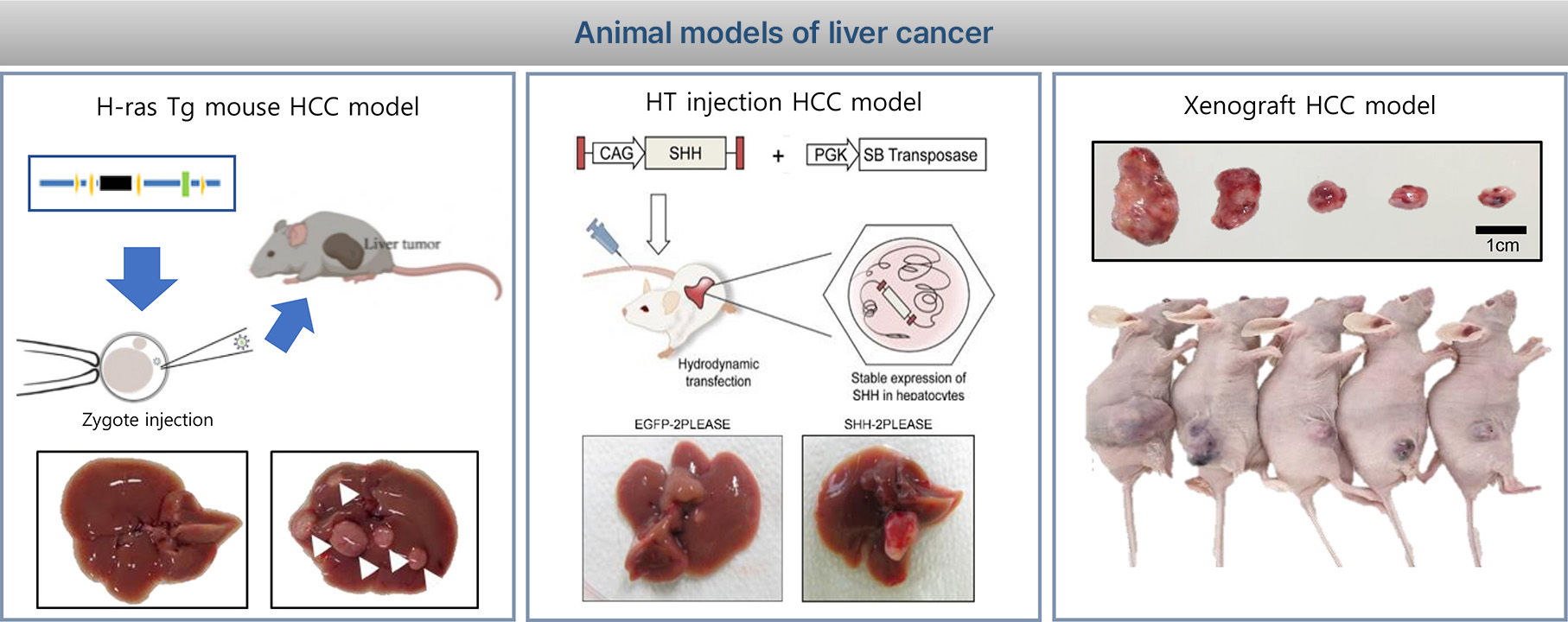
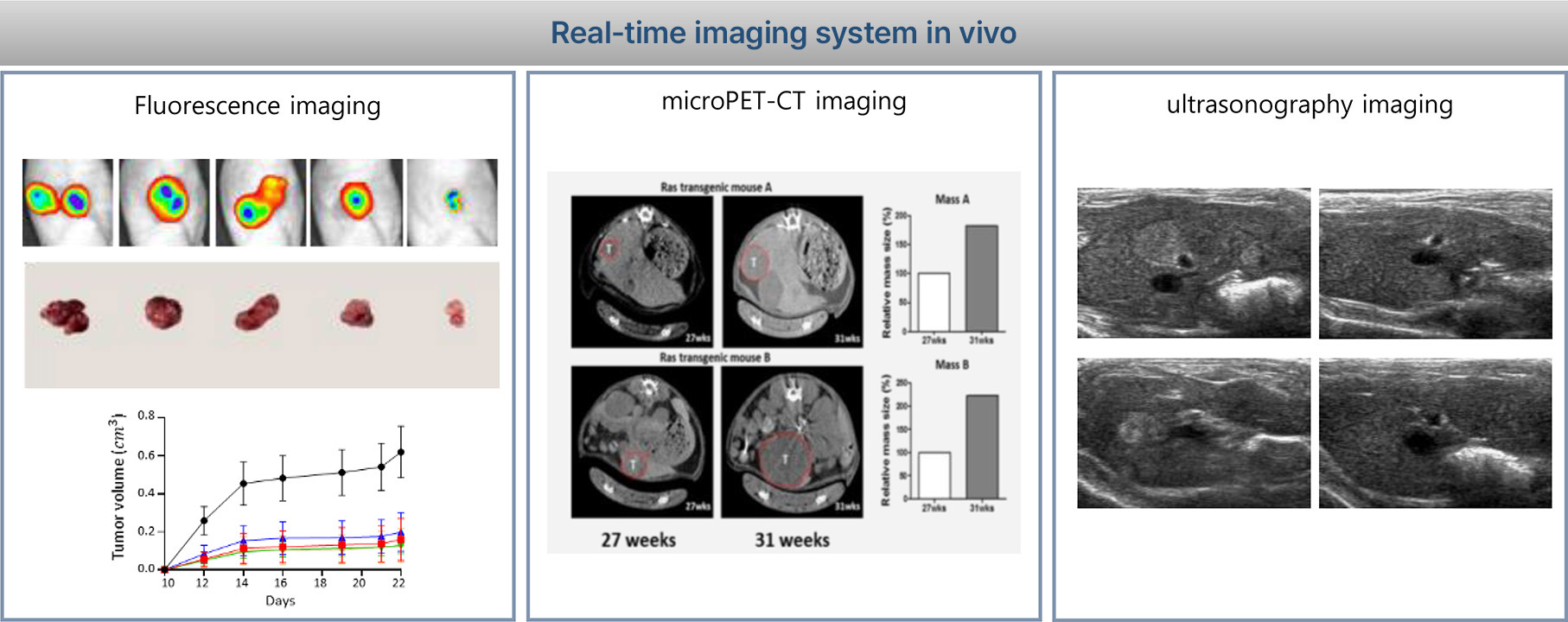
1. Liver cancer model
The H-ras12V transgenic (Ras-Tg) mice were originally developed by Dr. Dae-Yeoul Yu (Laboratory of Human Genomics, Korea Research Institute of Bioscience and Biotechnology, Daejeon, Korea). Male mice spontaneously developed HCC beginning at approximately 10-15 weeks of age. (Journal of Hepatology 43 (2015) 836-844). The Ras-Tg mouse liver cancer model is the most similar to human liver cancer. It is a unique liver cancer in vivo evaluation system that can assess liver cancer treatment effect by diagnosing initial liver cancer around 1mm using ultra-sonography and evaluating the effect of liver cancer with drug administration.
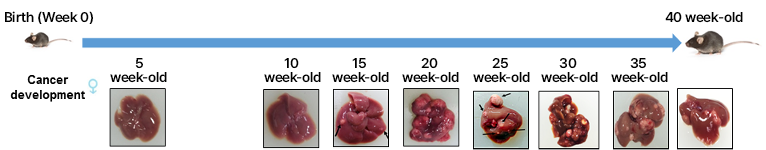
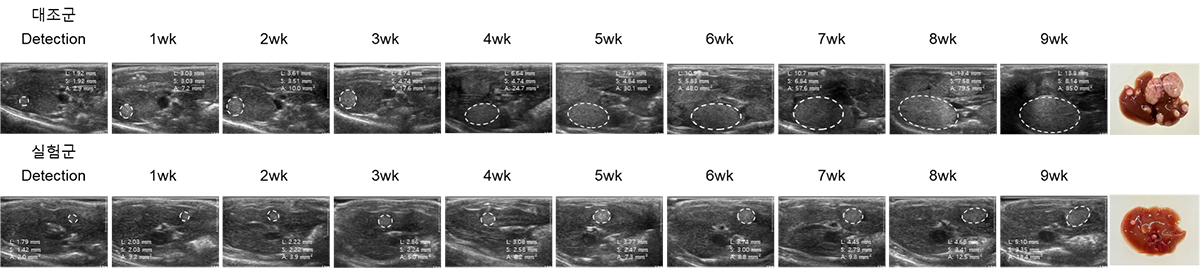
2. Lung cancer model
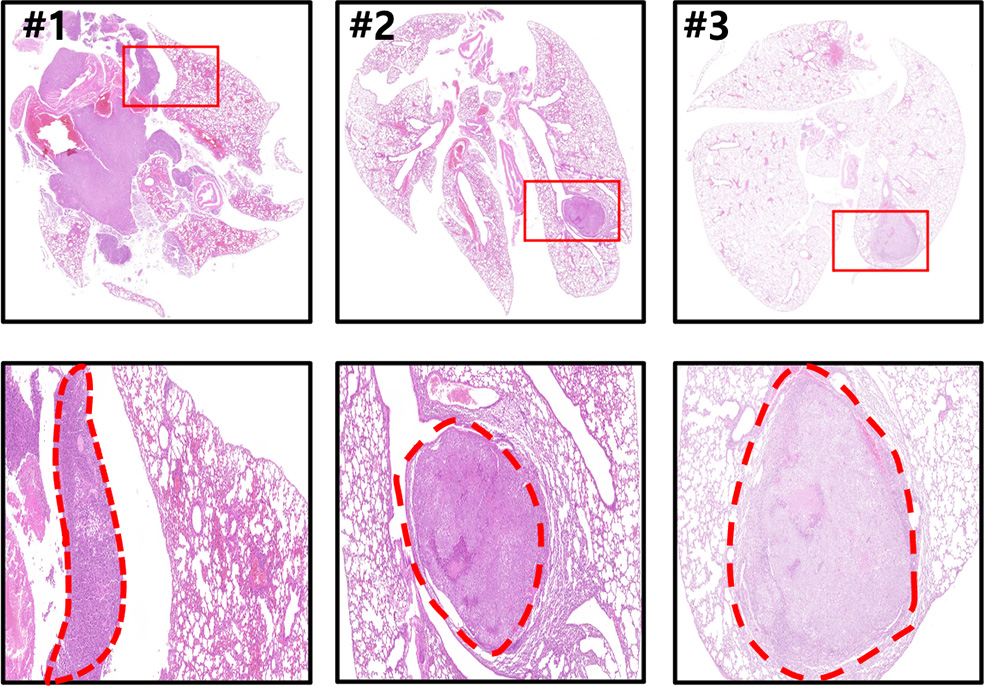
The established animal lung cancer model induces human lung cancer in the mouse lung by injecting a human lung cancer cell line to athymic nude mice into the tail vein. In contrast with the xenograft cancer animal model, it is possible to directly develop human cancer in the mouse lung to evaluate the therapeutic and tissue delivery effects of RNA drugs in lung cancer.
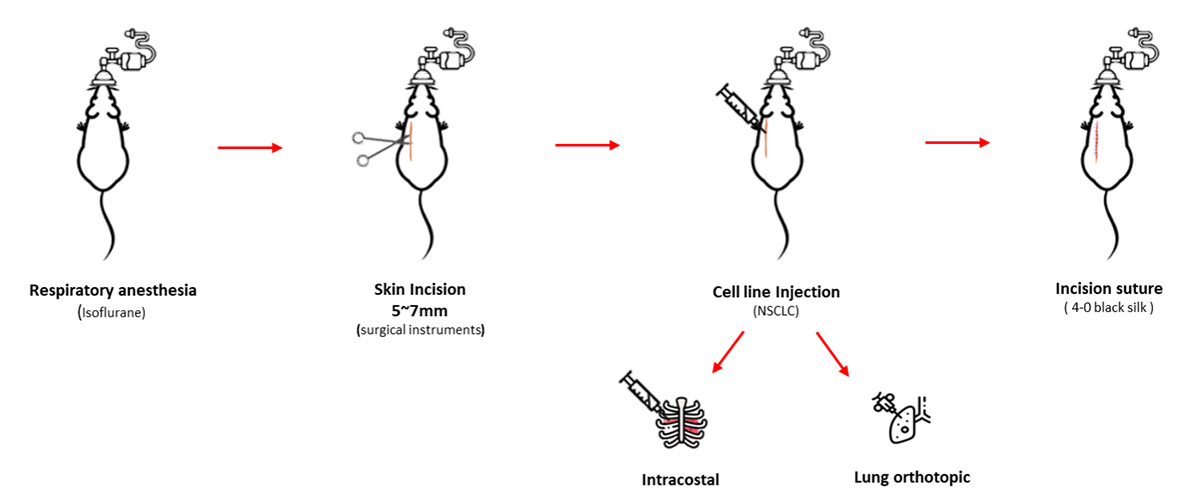
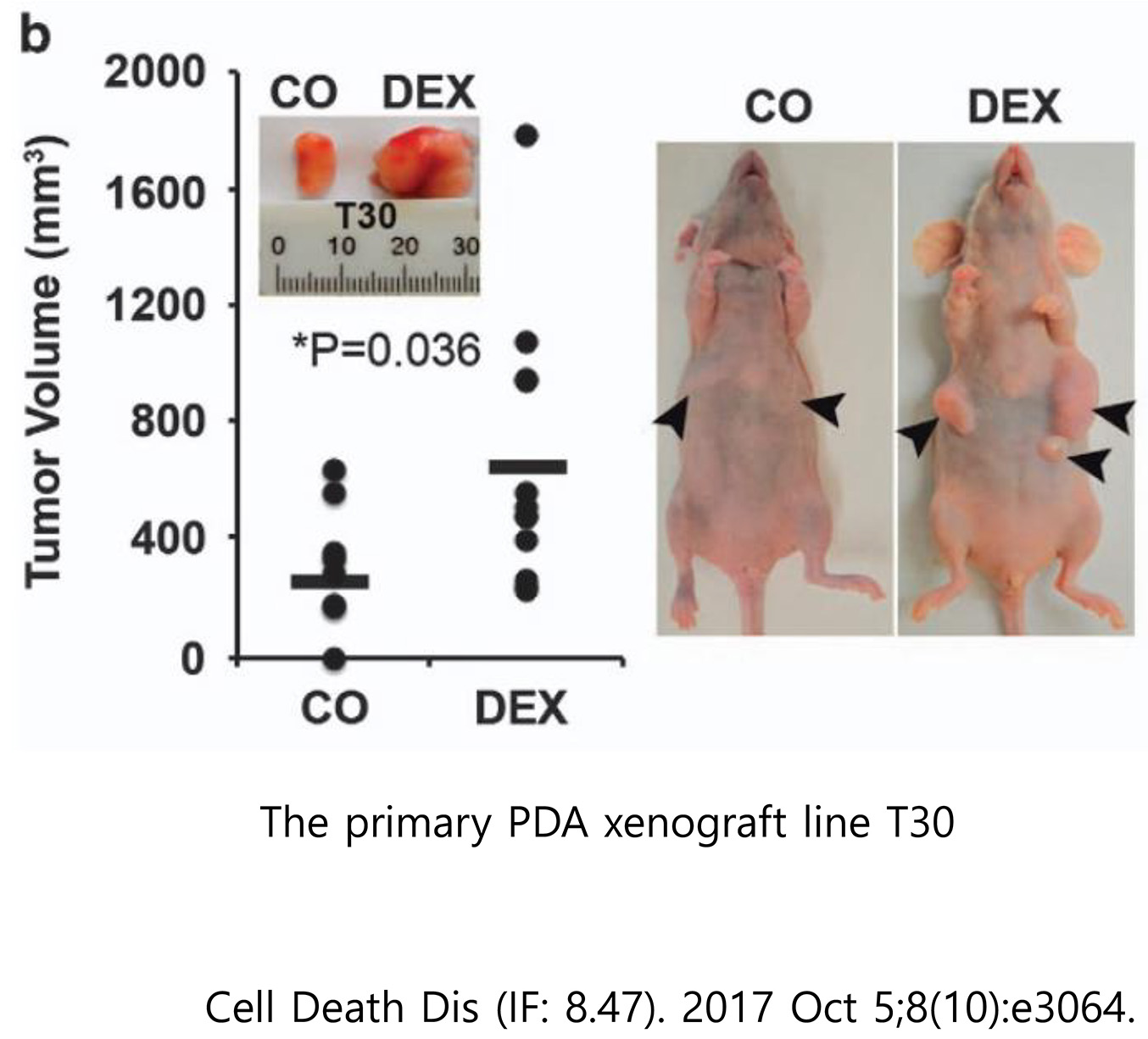
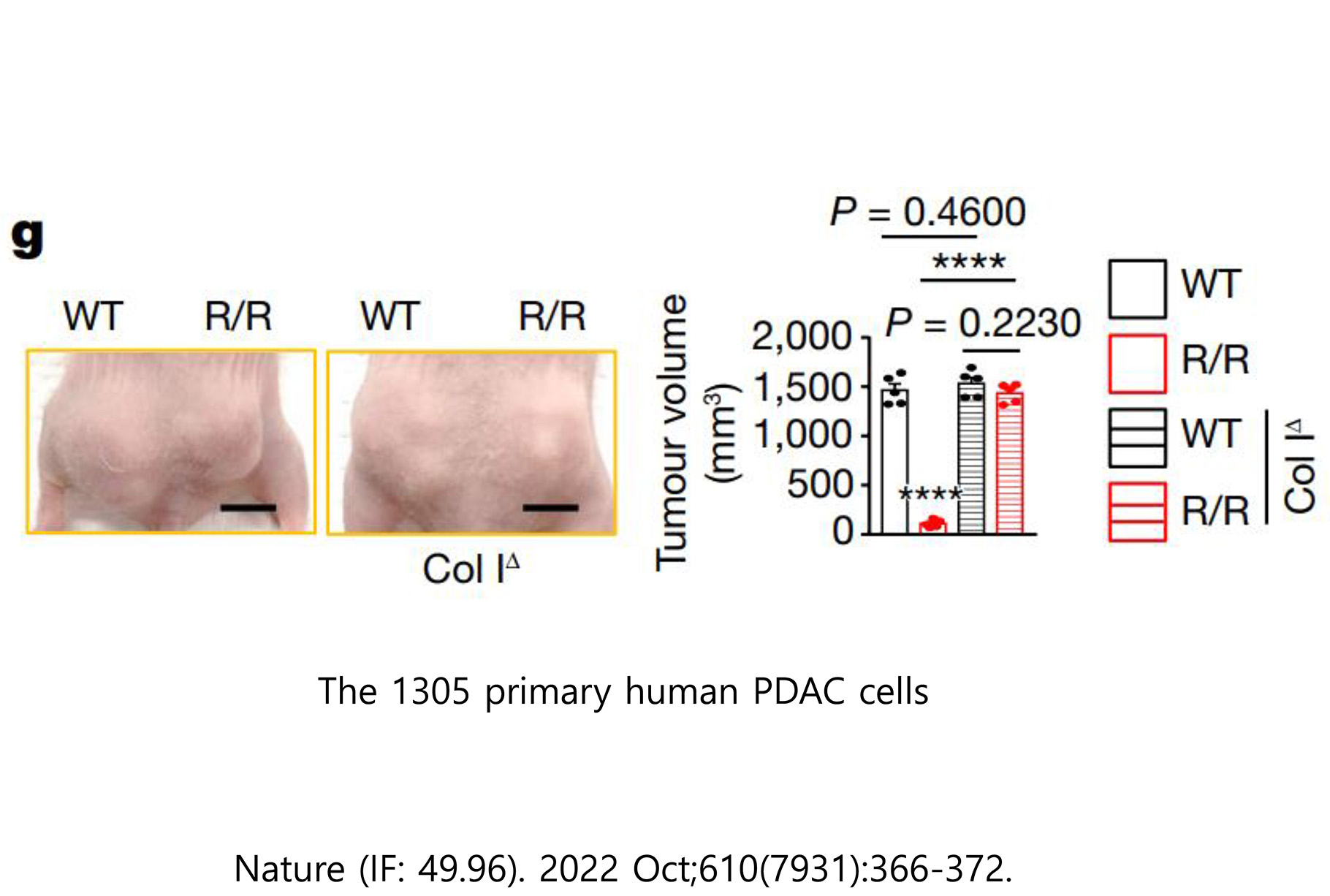
3. Pancreatic cancer model
The pancreatic cancer animal model uses a xenograft animal model of subcutaneous transplantation of human pancreatic cancer cell lines into athymic nude mice.
Tech. 3
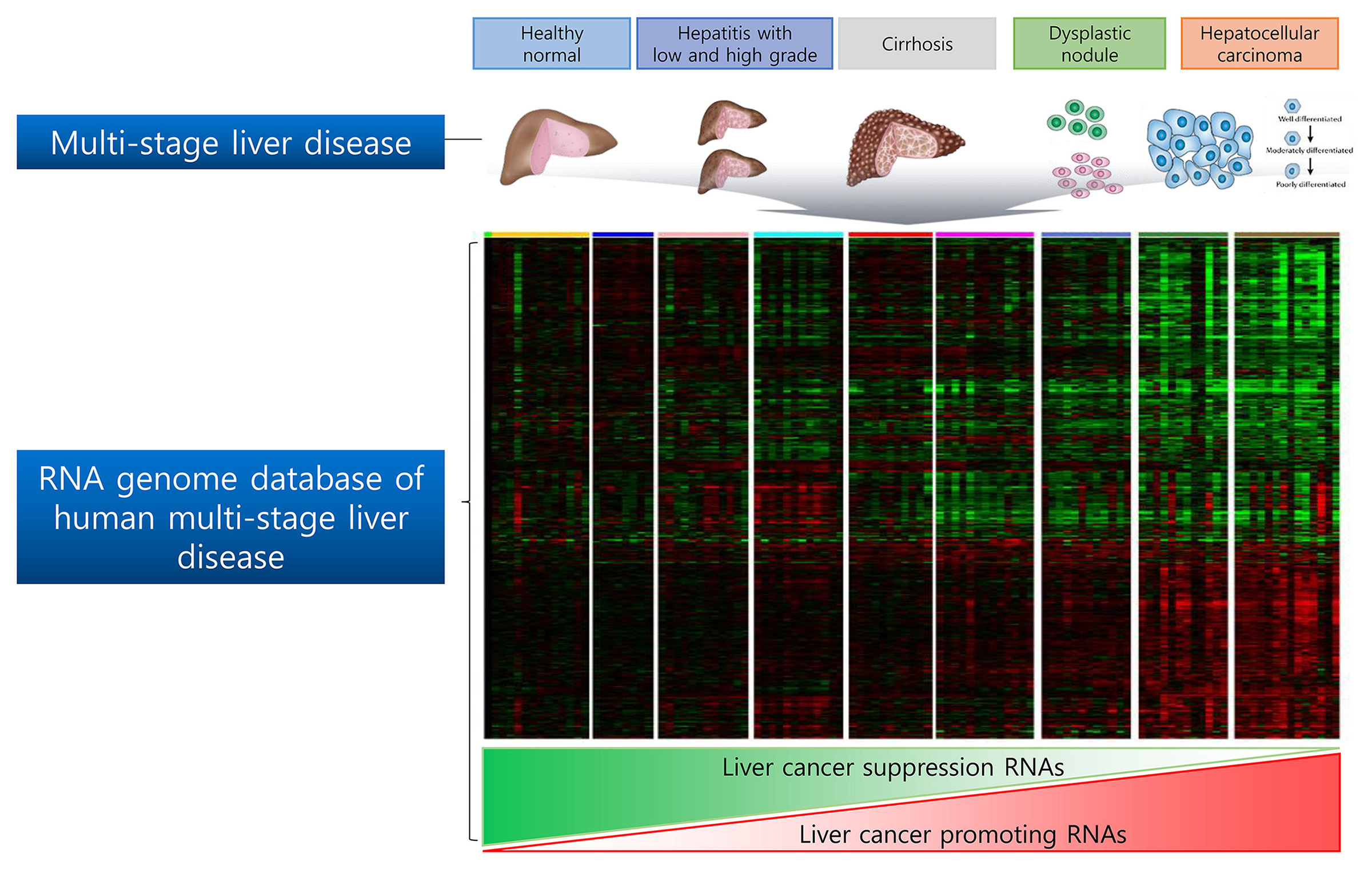
RNA Genome Database of Multi-stage Liver disease
NEORNAT Inc. has established RNA genome database (DB) of whole liver disease including healthy normal, hepatitis with low and high-grade fibrosis, cirrhosis, low and high-grade dysplastic nodule, and hepatocellular carcinoma (Edmonson Grade 1,2,3). This NEORNAT’s human liver disease RNA DB allow us to identify functional coding & non-coding RNAs contributing liver cancer progression and development. By performing in vitro and in vivo tumorigenesis analyses, these proof of concept candidates are then being validated for the first-in-class targets as developing pipeline for the non-clinical stage.
| Hardware | 4 high-performance big data analysis servers based on Linux |
|---|---|
| Software | Established a core pipeline for RNome analysis, and have data analysis tools such as BGW (Biomedical Genomics Workbench) |
| Open database | Genomic big data analysis of TCGA, ICGC, NCBI Gene Expression Omnibus, etc., RNA information database (cBioportal, COSMIC), miRNA target prediction database (TargetScan, miRWALK, miRGator, 4miRs), untranslated RNA function prediction database (starBase, InCeDB), miRcode, spongeScan |
